2006”N“x
ђXЉо‹а•сЌђЏ‘
Њ¤‹†‰Ы‘и–јЃF€в“`Ћq”Њ»‚Єѓ^ѓ“ѓpѓNЋїЉФ‘ЉЊЭЌм—p‚Й—^‚¦‚й‰e‹ї‚М‰рђН
Њcњд‹`Џm‘еЉw ђЌфҐѓЃѓfѓBѓAЊ¤‹†‰И ”ЋЋm‰Ы’ц1”N
–kђм“ќ”V
1.
Introduction
As phenotypes of organisms are not fully
described by only few sets of genes, we should take biochemical networks such
as Protein-Protein Interaction (PPI) and Protein-DNA Interaction (PDI) into
consideration. Recent progress of high-throughput PPI detection methods are
producing vast amount of PPI data in human (Rual
et al., 2005) as well as other model organisms. With the increase of data,
various tools and methods have been developed to analyze the data. However,
most of those methods are effective for only unicellular organisms. Higher
eukaryotes have highly differentiated cells and the gene expression patterns
vary greatly among the cell types. In addition, higher eukaryotes have a wide
variety of post-transcriptional modification, such as alternative splicing
(AS), polyadenylation and RNA editing. Among those modifications, AS would have
the largest impact on PPI networks (Figure 1).
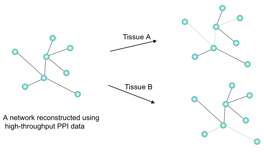
Figure 1.Ѓ@ A model of a protein interaction map in
higher eukaryotes. A reconstructed PPI network using high-throughput PPI data
cannot be used to retrieve biological meanings because highly differentiated
cells have different gene expression and post-transcriptional modification.
More than half of human genes thought to be
alternatively spliced. That means many PPI motifs might be alternatively used
as shown in Figure 2. There is a research which demonstrated alternative
splicing are statistically not related to the border of protein motifs (Offman
et al., 2004). The result of the research seems to suggest AS is not important
for PPI networks. However, the result also demonstrated AS events do not avoid
PPI motifs, suggesting AS could change PPI networks greatly if different AS
variants are expressed in different types of cells. As a lot of gene expression
data support this assumption, I decided to investigate the impact of AS on PPI
networks concretely.
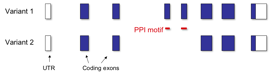

Figure 2.Ѓ@ A model of a gene which has alternatively
deleted PPI motif. This gene would not interact other proteins through the PPI
motif when the variant 2 are expressed dominantly.
2.
Materials and Methods
I used Ensembl Release 40 (Birney
et al., 2006) as MySQL data, especially human gene expression (EST) and protein
features (signal peptide, transmembrane, low complexity, coil, InterPro) in
BioMart. These data were downloaded from ftp.ensembl.org in September 2006.
Table 1.Ѓ@ Materials (Ensembl release-40 coding gene
and transcript data)
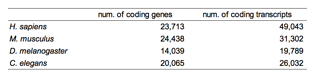
Although Ensembl registered genes are more
than described in Table 1 (e.g., 31,718 genes and 57,048 transcripts in human
data), I used coding genes as parent population in this study because the
objective are investigating alternative protein motifs.
I also used protein descriptions of
InterPro (Mulder
et al., 2007), pathway data of KEGG (Kanehisa
et al., 2006) through KEGG API, and AEdb, manual generated alternative splicing
data in ASD (Stamm
et al., 2006).
In Ensembl protein features, coiled coil
regions are annotated with ncoils program (Lupas
et al., 1991), low-complexity regions are annotated with SEG program (Wootton
and Federhen, 1996),
signal sequence regions are characterised with SignalP (Nielsen
et al., 1997; Nielsen and Krogh, 1998; Bendtsen
et al., 2004), and transmembrane regions are annotated with TMHMM. (Sonnhammer
et al., 1998; Krogh et al., 2001)
3.
Results and Discussion
A few information about experimentally
demonstrated alternative splicing events that would change biochemical
reactions
To investigate how much experimentally
demonstrated alternative motifs exist, I searched ЃgRegulary featureЃh in
sequence data file of manual generated AS database AEdb. As a result, 55.4% of
all entries (2244) were unclear. ЃgStop codonЃh and ЃgFrameshiftЃh were 11.6% and
6.7%, respectively. The entries including ЃgbindЃh in the other 26.3% were only
64 (2.9%). This means binding regulatory AS is small ratio as Offman et al. suggest, or it is difficult to detect the binding function of AS
event experimentally. In the 64 entries, human, mouse, Norway rat were 23, 20,
9 entries, respectively. This result suggests it is important to analyze
genome-wide transcript data from the perspective of alternative motifs.
Higher eukaryotes tend to have more
alternative motif genes
To investigate genome-wide alternative
protein motifs and the difference among model eukaryotes (H. sapiens, M.
musculus, D. melanogaster and C. elegans), I used common Ensembl protein features (coiled coil, signal
peptide, transmembrane, low complexity). The results are shown in Table 2.
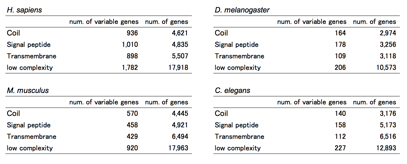
Table 2.Ѓ@ The number of genes which have variable
motifs. ЃgVariable genesЃh means the genes which have the motif when expressed as
specific variants of the alternative splicing events.
The ratios of the number of variable genes
to the number of genes are shown in Figure 3. This result suggests higher
eukaryotes tend to have more alternative motif genes. That also suggests that
the variety of phenotypes of higher eukaryotes are caused by alternative
motifs, considering the number of human genes are fewer than that of predicted.
As shown in Figure 3, low complexity motifs are likely to have less important
protein motifs than the other protein features. In addition, low complexity
regions could be found in many genes for their weak consensus, suggesting low
complexity region might be more highly estimated than they are.
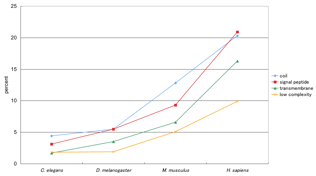
Figure 3.Ѓ@ The rate of genes which have variable motifs
among the transcripts.
To confirm that this result was not
influenced by the number of transcripts data for each species, I calculated the
rate of transcripts for each coding gene which is registered in Ensembl (Figure
4). As shown in Figure 4, the rate of M. musculus transcripts are fewer than that of D. melanogaster, and the rate of C. elegans
transcripts are nearly equal to that of M. musculus transcripts. These results suggest the result of Figure 3 is not
strongly influenced by the number of transcripts.
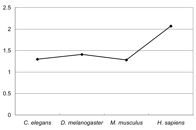
Figure 4.Ѓ@ The rate of transcripts for each coding gene
which are registered in Ensembl.
Types of the most influenced protein
motifs by alternative splicing in human
By the analyses of common protein motifs,
it was suggested that the phenotypes of higher eukaryotes could be reflected
the wide variety of alternative motifs. Then I focused on the human alternative
motifs to investigate what types of motifs are deleted in alternative splicing
events.
I investigated the most variable InterPro
motifs, which are deleted in a specific alternative splicing variants (Table
3), and the most variable binding related InterPro motifs for investigating the
changeability of PPI networks (Table 4).
Ѓ@Ѓ@Ѓ@Ѓ@Ѓ@ Table 3.Ѓ@ Variable
InterPro motifsЃ@Ѓ@Ѓ@Ѓ@Ѓ@Ѓ@Ѓ@Ѓ@Ѓ@Ѓ@ Table 4.Ѓ@ Variable InterPro binding motifs

The result of Table 3 suggests variable
alternative motifs are not restricted to PPI motifs and various types of
biochemical reactions could be changed through AS events. Most of Ѓgzinc fingerЃh
works as DNA binding motif, which support IVVЃfs c-Fos variants result. The C2H2
is highly counted probably because it is well-explored class. Other types of
zinc finger are also found (not shown in Table 3). Many types of kinase region
(e.g. Wall-associated kinase) are found, which suggests AS can change signaling
cascades without PPI changes. ЃgToropmyosinЃh variants can vary cytoskeleton
through actin-binding, which is also supported by the result of Table 4. Pleckstrin
homology domain (PH domain) is also thought to participate in signaling
cascades. The function of ЃgEGF-like regionЃh is not yet clear. However, the
motif is found in extracellular domain and there is a research which report
that ЃgImmunoglobulin-likeЃh domain is probably essential for efficient
interaction of an EGF-like domain with ErbB receptors (Eto
et al., 2006). The number of ЃgProline-rich regionЃh possibly reflects its low
specificity. These results suggest alternative motifs could regulate signaling
cascades via various reactions such as PPI, PDI, receptor-ligand interaction,
kinase activity, and cytoskeletal changes.
Alternative Ca2+ binding
motifs and their expression patters
Table 4 shows the result of binding-related
alternative motifs. This result is consistent with that of Table 3. As the result
suggest the importance of Ca2+ binding, I conducted further
investigation about alternative motifs which can change Ca2+
binding.
To investigate the importance of each
alternative Ca2+ binding gene in signaling cascades, I mapped the
genes which would change Ca2+ binding to KEGG pathways by KEGG API.
An example of the results is shown in Figure 5. The path which connect the
genes and Ca2+ would be cut when the alternative variants are
dominantly expressed.
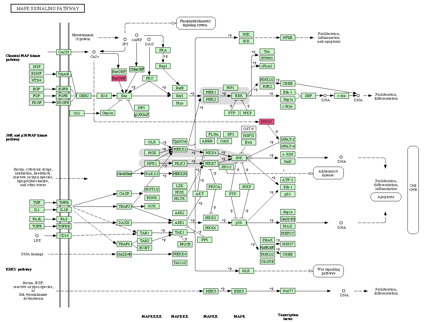
Figure 5.Ѓ@ An example of the pathways including genes which have variable calcium binding motifs. Ras signaling and/or NFAT regulated gene expression would change in MAPK signaling cascades when the motifs are not in dominantly expressed AS variants.
AS variants which do not have
calcium-binding motif could be aberrant variants. Therefore, I examined the
tissues in which AS variants having calcium-binding motif or not. As a result,
only Calcineurin subunit B gene (Ensembl gene ID: ENSG00000115953) has
different expressed tissue pattern. In pathological data, AS variants which do
not have calcium-binding motif are expressed at 'retinoblastoma', 'leukemia',
'carcinoid', 'ascites' and 'lymphoblastic'. These are not described in OMIM (601302),
suggesting the AS variants cause those diseases via changing signaling
cascades.
Changing protein motif via AS might be a
cause of neural diversity in human
To investigate whether motif changing AS
events are important for diversity of the human phenotype, I examined the rate
of genes which have variable motifs (signal peptide, coiled coil,
transmembrane, low complexity) to genes which do not have variable motifs. The
result is shown in Table 5. Many neural cells related tissues are ranked in
top10, especially in signal peptide. That might not reflect the bias of the
original dataset because low complexity motifs are thought to be less important
because of its weak consensus and neural cell related tissues are not highly
ranked in low complexity.
Table 5.Ѓ@ The rate of genes which have alternative motifs and expressed in each tissue (anatomy). The rate was calculated as the ratio of alternative motif genes to non-alternative motif genes. Red-colored anatomies are neural cell related.
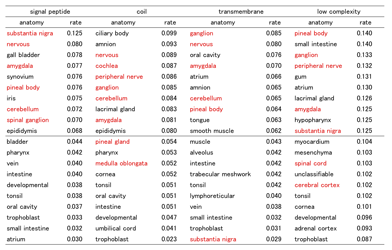
Until now, some research has tried to link the diversity of human phenotype and post-transcriptional modification especially AS, however strong evidence was not shown in those researches. Taking account of this study, motif changing AS would be important rather than AS events itself, also suggesting not all AS variants are biologically essential.
References
Bendtsen, J.D., Nielsen, H., von Heijne, G.
and Brunak, S. (2004) J Mol Biol, 340, 783-95.
Birney, E.,
Andrews, D., Caccamo, M., Chen, Y., Clarke, L., Coates, G., Cox, T.,
Cunningham, F., Curwen, V., Cutts, T., et al. (2006) Nucleic Acids Res, 34, D556-61.
Eto, K., Eda, K.,
Kanemoto, S. and Abe, S. (2006) Biochem Biophys Res Commun, 350, 263-71.
Kanehisa, M.,
Goto, S., Hattori, M., Aoki-Kinoshita, K.F., Itoh, M., Kawashima, S., Katayama,
T., Araki, M. and Hirakawa, M. (2006) Nucleic Acids Res, 34, D354-7.
Krogh, A.,
Larsson, B., von Heijne, G. and Sonnhammer, E.L. (2001) J Mol Biol, 305, 567-80.
Lupas, A., Van
Dyke, M. and Stock, J. (1991) Science, 252, 1162-4.
Mulder, N.J.,
Apweiler, R., Attwood, T.K., Bairoch, A., Bateman, A., Binns, D., Bork, P.,
Buillard, V., Cerutti, L., Copley, R., et al. (2007) Nucleic Acids Res, 35, D224-8.
Nielsen, H.,
Engelbrecht, J., Brunak, S. and von Heijne, G. (1997) Protein Eng, 10, 1-6.
Nielsen, H. and
Krogh, A. (1998) Proc Int Conf Intell Syst Mol Biol, 6, 122-30.
Offman, M.N.,
Nurtdinov, R.N., Gelfand, M.S. and Frishman, D. (2004) BMC Bioinformatics, 5, 41.
Rual, J.F.,
Venkatesan, K., Hao, T., Hirozane-Kishikawa, T., Dricot, A., Li, N., Berriz,
G.F., Gibbons, F.D., Dreze, M., Ayivi-Guedehoussou, N., et al. (2005) Nature, 437, 1173-8.
Sonnhammer, E.L.,
von Heijne, G. and Krogh, A. (1998) Proc Int Conf Intell Syst Mol Biol, 6, 175-82.
Stamm, S.,
Riethoven, J.J., Le Texier, V., Gopalakrishnan, C., Kumanduri, V., Tang, Y.,
Barbosa-Morais, N.L. and Thanaraj, T.A. (2006) Nucleic Acids Res, 34, D46-55.
Wootton, J.C. and
Federhen, S. (1996) Methods Enzymol, 266, 554-71.